Univ.-Prof. Dr. David Berry

Research in David Berry’s group is focused on understanding the biology of the human microbiota and its role in health and disease. He has pioneered the development of novel experimental and computational tools to reveal the functionof microbial communities and has developed single cell isotope labelling techniques to characterise functional guilds in the intestinal ecosystem.
The main research aims of David’s group are to gain a fundamental understanding of the assembly and interactions of the intestinal microbiota and to uncover how the microbiota affects host physiology. David is active in translational and clinical research in several fields, including chronic inflammation, nutrition, metabolic syndrome, cancer, and gut-immune-brain axis development in newborns. He is coordinator of an Austrian Science Fund (FWF) research consortium focused on premature infant health (2023).
Recent projects
Deciphering microbial interactions and detecting keystone species with co-occurrence networks.
Co-occurrence networks produced from microbial survey sequencing data are frequently used to identify interactions between community members, thoughmany questions remain about their validity and performance. Using a simulation-based approach, we evaluated how well networks reveal underlying interactions. We found that co-occurrence networks can recapitulate interaction networks under certain conditions, and also identified topological features associated with keystone species in co-occurrence networks. This work provides a substantiated framework to guide environmental microbiologists in the construction and interpretation of co-occurrence networks from microbial survey datasets. (Berry and Widder, 2014)
Single cell stable isotope probing of the gut microbiota
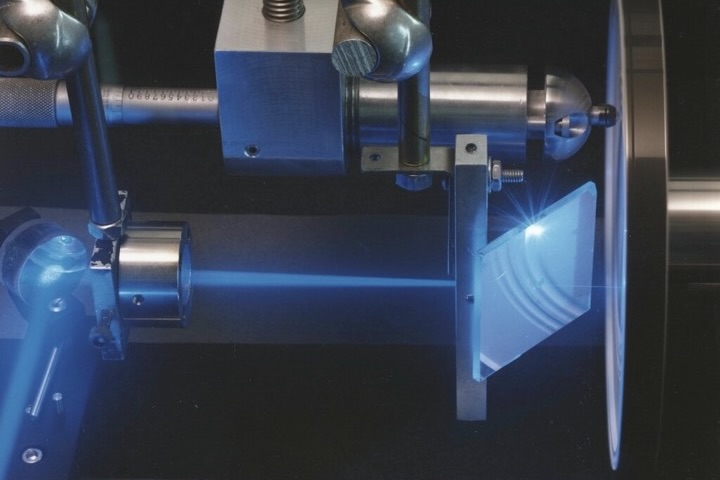
Microbes live almost exclusively in complex communities, and it remains a major challenge to identify the activity of microbial cells under natural conditions. We have developed tools based on single cell stable isotope probing to address this. Using fluorescence in situ hybridization combined with high-resolution secondary ion mass spectrometry (NanoSIMS) we were able to identify microbial cells that had degraded secreted mucosal proteins (Berry et al., 2013). Recently, we were able to use heavy water as a universal activity marker combined with Raman microspectroscopy to profile the in situ activity of two mucus-degrading species, Akkermansia muciniphila and Bacteroides acidifaciens, to a range of amended nutrients (Berry et al., 2015).
Microbiota and intestinal inflammation
Human inflammatory bowel diseases are thought to be affected by the gut microbiota and are characterized by an altered gut microbiota composition (Berry and Reinisch, 2013). We have been investigating this relationship using mouse colitis models (Berry et al., 2012; Schwab et al., 2014) as well as by evaluating the potential of fecal microbiota transplantation for the treatment of ulcerative colitis in humans (Angelberger et al., 2013).
Research Projects
Group Members
- Franziska Bauchinger
- Viktoria Maria Berndorfer
- Anna-Celine Danklmaier
- Kirsten Gassner
- Dr. Adnan Hodžić
- Mina Joseph
- Mia Juracic
- Sanaz Khadem
- Dr. Michaela Lang
- Georgi Nikolov
- Miriam Philipp
- Hamid Rasoulimehrabani
- Franka Schneider
- Dr. Stephanie Schnorr
- Dinely Wijayagunasekera
- Jan-Philipp Wittlinger
- Christos Zioutis
- Ekaterina Zolotoreva